(The below post contains excerpts from Slithering your way into bioinformatics with snakemake, Hackmd Press, 2023.)
Installation and setup!
Setup and installation
I suggest working in a new directory.
You'll need to install snakemake and sourmash. We suggest using mamba, via miniforge/mambaforge, for this.
Getting the data:
You'll need to download these three files:
- GCF_000021665.1_ASM2166v1_genomic.fna.gz
- GCF_000017325.1_ASM1732v1_genomic.fna.gz
- GCF_000020225.1_ASM2022v1_genomic.fna.gz
and rename them so that they are in a subdirectory genomes/ with the names:
GCF_000017325.1.fna.gz
GCF_000020225.1.fna.gz
GCF_000021665.1.fna.gz
Note, you can download saved copies of them here, with the right names: osf.io/2g4dm/.
Chapter 1 - snakemake runs programs for you!
Bioinformatics often involves running many different programs to characterize and reduce sequencing data, and I use snakemake to help me do that.
A first, simple snakemake workflow
Here's a simple, useful snakemake workflow:
rule compare_genomes:
message: "compare all input genomes using sourmash"
shell: """
sourmash sketch dna -p k=31 genomes/*.fna.gz --name-from-first
sourmash compare GCF_000021665.1.fna.gz.sig \
GCF_000017325.1.fna.gz.sig GCF_000020225.1.fna.gz.sig \
-o compare.mat
sourmash plot compare.mat
"""
Put it in a file called Snakefile, and run it with snakemake -j 1.
This will produce the output file compare.mat.matrix.png which contains a similarity matrix and a dendrogram of the three genomes (see Figure 1).
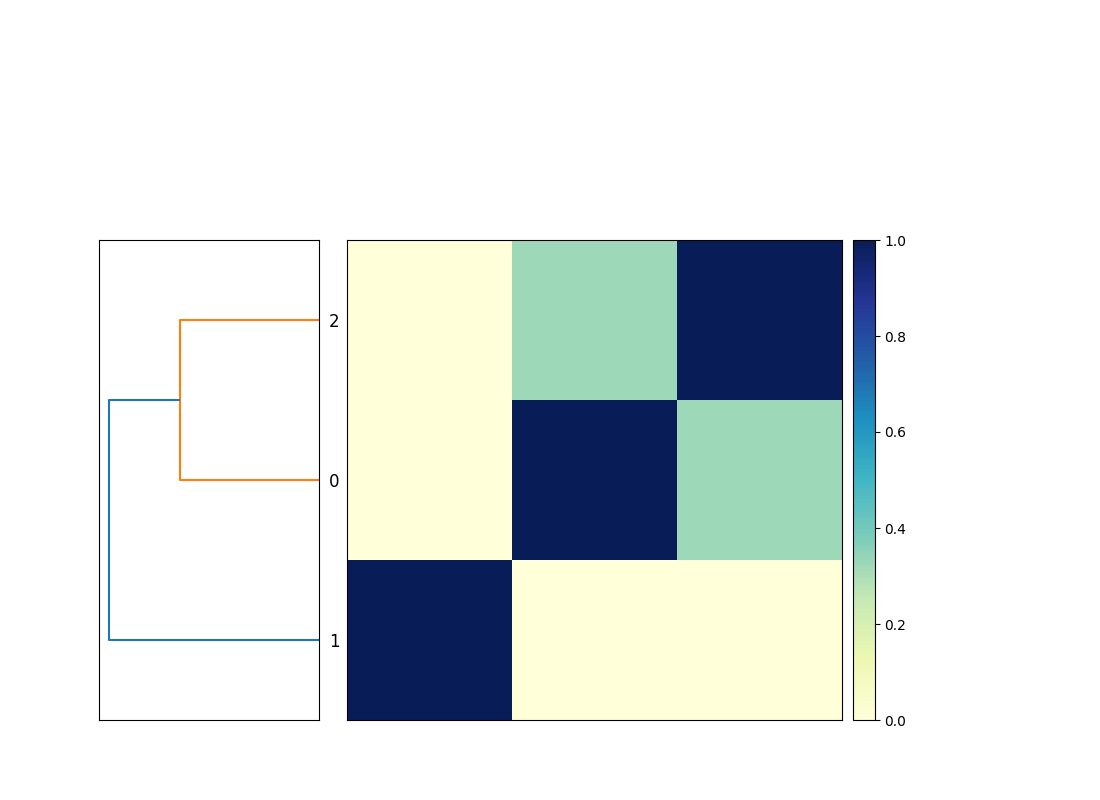
This is functionally equivalent to putting these three commands into a file compare-genomes.sh and running it with bash compare-genomes.sh -
sourmash sketch dna -p k=31 genomes/*.fna.gz --name-from-first
sourmash compare GCF_000021665.1.fna.gz.sig \
GCF_000017325.1.fna.gz.sig GCF_000020225.1.fna.gz.sig \
-o compare.mat
sourmash plot compare.mat
The snakemake version is already a little bit nicer because it will give you encouragement when the commands run successfully (with nice green text saying "1 of 1 steps (100%) done"!) and if the commands fail you'll get red text alerting you to that, too.
But! We can further improve the snakemake version over the shell script version!
Avoiding unnecessary rerunning of commands: a second snakemake workflow
The commands will run every time you invoke snakemake with snakemake -j 1. But most of the time you don't need to rerun them because you've already got the output files you wanted!
How do you get snakemake to avoid rerunning rules?
We can do that by telling snakemake what we expect the output to be by adding an output: block in front of the shell block:
rule compare_genomes:
message: "compare all input genomes using sourmash"
output:
"compare.mat.matrix.png"
shell: """
sourmash sketch dna -p k=31 genomes/*.fna.gz --name-from-first
sourmash compare GCF_000021665.1.fna.gz.sig \
GCF_000017325.1.fna.gz.sig GCF_000020225.1.fna.gz.sig \
-o compare.mat
sourmash plot compare.mat
"""
and now when we run snakemake -j 1 once, it will run the commands; but when we run it again, it will say, "Nothing to be done (all requested files are present and up to date)."
This is because the desired output file, compare.mat.matrix.png, already exists. So snakemake knows it doesn't need to do anything!
If you remove compare.mat.matrix.png and run snakemake -j 1 again, snakemake will happily make the files again:
rm compare.mat.matrix.png
snakemake -j 1
So snakemake makes it easy to avoid re-running a set of commands if it has already produced the files you wanted. This is one of the best reasons to use a workflow system like snakemake for running bioinformatics workflows; shell scripts don't automatically avoid re-running commands.
Running only the commands you need to run
The last Snakefile above has three commands in it, but if you remove the compare.mat.matrix.png file you only need to run the last command again - the files created by the first two commands already exist and don't need to be recreated. However, snakemake doesn't know that - it treats the entire rule as one rule, and doesn't look into the shell command to work out what it doesn't need to run.
If we want to avoid re-creating the files that already exist, we need to make the Snakefile a little bit more complicated.
First, let's break out the commands into three separate rules.
rule sketch_genomes:
shell: """
sourmash sketch dna -p k=31 genomes/*.fna.gz --name-from-first
"""
rule compare_genomes:
shell: """
sourmash compare GCF_000021665.1.fna.gz.sig \
GCF_000017325.1.fna.gz.sig GCF_000020225.1.fna.gz.sig \
-o compare.mat
"""
rule plot_comparison:
message: "compare all input genomes using sourmash"
output:
"compare.mat.matrix.png"
shell: """
sourmash plot compare.mat
"""
We didn't do anything too complicated here - we made two new rule blocks, with their own names, and split the shell commands up so that each shell command has its own rule block.
You can tell snakemake to run all three:
snakemake -j 1 sketch_genomes compare_genomes plot_comparison
and it will successfully run them all!
However, we're back to snakemake running some of the commands every time - it won't run plot_comparison every time, because compare.mat.matrix.png exists, but it will run sketch_genomes and compare_genomes repeatedly.
How do we fix this?
Adding output blocks to each rule
If add output blocks to each rule, then snakemake will only run rules where the output needs to be updated (e.g. because it doesn't exist).
Let's do that:
rule sketch_genomes:
output:
"GCF_000017325.1.fna.gz.sig",
"GCF_000020225.1.fna.gz.sig",
"GCF_000021665.1.fna.gz.sig"
shell: """
sourmash sketch dna -p k=31 genomes/*.fna.gz --name-from-first
"""
rule compare_genomes:
output:
"compare.mat"
shell: """
sourmash compare GCF_000021665.1.fna.gz.sig \
GCF_000017325.1.fna.gz.sig GCF_000020225.1.fna.gz.sig \
-o compare.mat
"""
rule plot_comparison:
message: "compare all input genomes using sourmash"
output:
"compare.mat.matrix.png"
shell: """
sourmash plot compare.mat
"""
and now
snakemake -j 1 sketch_genomes compare_genomes plot_comparison
will run each command only once, as long as the output files are still there. Huzzah!
But we still have to specify the names of all three rules, in the right order, to run this. That's annoying! Let's fix that next.
Chapter 2: snakemake connects rules for you!
Chaining rules with input: blocks
We can get snakemake to automatically connect rules by providing information about the input files a rule needs. Then, if you ask snakemake to run a rule that requires certain inputs, it will automatically figure out which rules produce those inputs as their output, and automatically run them.
Let's add input information to the plot_comparison and compare_genomes
rules:
rule sketch_genomes:
output:
"GCF_000017325.1.fna.gz.sig",
"GCF_000020225.1.fna.gz.sig",
"GCF_000021665.1.fna.gz.sig"
shell: """
sourmash sketch dna -p k=31 genomes/*.fna.gz --name-from-first
"""
rule compare_genomes:
input:
"GCF_000017325.1.fna.gz.sig",
"GCF_000020225.1.fna.gz.sig",
"GCF_000021665.1.fna.gz.sig"
output:
"compare.mat"
shell: """
sourmash compare GCF_000021665.1.fna.gz.sig \
GCF_000017325.1.fna.gz.sig GCF_000020225.1.fna.gz.sig \
-o compare.mat
"""
rule plot_comparison:
message: "compare all input genomes using sourmash"
input:
"compare.mat"
output:
"compare.mat.matrix.png"
shell: """
sourmash plot compare.mat
"""
Now you can just ask snakemake to run the last rule:
snakemake -j 1 plot_comparison
and snakemake will run the other rules only if those input files don't exist and need to be created.
Taking a step back
The Snakefile is now a lot longer, but it's not too much more complicated - what we've done is split the shell commands up into separate rules and annotated each rule with information about what file it produces (the output), and what files it requires in order to run (the input).
This has the advantage of making it so you don't need to rerun commands unnecessarily. This is only a small advantage with our current workflow, because sourmash is pretty fast. But if each step takes an hour to run, avoiding unnecessary steps can make your work go much faster!
And, as you'll see later, these rules are reusable building blocks that can be incorporated into workflows that each produce different files. So there are other good reasons to break shell commands out into individual rules!
Chapter 3: snakemake helps you avoid redundancy!
Avoiding repeated filenames by using {input} and {output}
If you look at the previous Snakefile, you'll see a few repeated filenames - in particular, rule compare_genomes has three filenames in the input block and then repeats them in the shell block, and compare.mat is repeated several times in both compare_genomes and plot_genomes.
We can tell snakemake to reuse filenames by using {input} and {output}. The { and } tell snakemake to interpret these not as literal strings but as template variables that should be replaced with the value of input and output.
Let's give it a try!
rule sketch_genomes:
output:
"GCF_000017325.1.fna.gz.sig",
"GCF_000020225.1.fna.gz.sig",
"GCF_000021665.1.fna.gz.sig"
shell: """
sourmash sketch dna -p k=31 genomes/*.fna.gz --name-from-first
"""
rule compare_genomes:
input:
"GCF_000017325.1.fna.gz.sig",
"GCF_000020225.1.fna.gz.sig",
"GCF_000021665.1.fna.gz.sig"
output:
"compare.mat"
shell: """
sourmash compare {input} -o {output}
"""
rule plot_comparison:
message: "compare all input genomes using sourmash"
input:
"compare.mat"
output:
"compare.mat.matrix.png"
shell: """
sourmash plot {input}
"""
This approach not only involves less typing in the first place, but also makes it so that you only have to edit filenames in one place. This avoids mistakes caused by adding or changing filenames in one place and not another place - a mistake I've made plenty of times!
snakemake makes it easy to rerun workflows!
It is common to want to rerun an entire workflow from scratch, to make sure that you're using the latest data files and software. Snakemake makes this easy!
You can ask snakemake to clean up all the files that it knows how to generate - and only those files:
snakemake -j 1 plot_comparison --delete-all-output
which can then be followed by asking snakemake to regenerate the results:
snakemake -j 1 plot_comparison
snakemake will alert you to missing files if it can't make them!
Suppose you add a new file that does not exist to compare_genomes:
rule sketch_genomes:
output:
"GCF_000017325.1.fna.gz.sig",
"GCF_000020225.1.fna.gz.sig",
"GCF_000021665.1.fna.gz.sig"
shell: """
sourmash sketch dna -p k=31 genomes/*.fna.gz --name-from-first
"""
rule compare_genomes:
input:
"GCF_000017325.1.fna.gz.sig",
"GCF_000020225.1.fna.gz.sig",
"GCF_000021665.1.fna.gz.sig",
"does-not-exist.sig"
output:
"compare.mat"
shell: """
sourmash compare {input} GCF_000021665.1.sig -o {output}
"""
rule plot_comparison:
message: "compare all input genomes using sourmash"
input:
"compare.mat"
output:
"compare.mat.matrix.png"
shell: """
sourmash plot {input}
"""
Here, does-not-exist.sig doesn't exist, and we haven't given snakemake a rule to make it, either. What will snakemake do??
It will complain, loudly and clearly! And it will do so before running anything.
First, let's force the rule remove the output file that depends on the
rm compare.mat
and then run snakemake -j 1. You should see:
Missing input files for rule compare_genomes:
output: compare.mat
affected files:
does-not-exist.sig
This is exactly what you want - a clear indication of what is missing before your workflow runs.
Next steps
We've introduced basic snakemake workflows, which give you a simple way to run shell commands in the right order. snakemake already offers a few nice improvements over running the shell commands by yourself or in a shell script -
- it doesn't run shell commands if you already have all the files you need
- it lets you avoid typing the same filenames over and over again
- it gives simple, clear errors when something fails
While this functionality is nice, there are many more things we can do to improve the efficiency of our bioinformatics!
In the next section, we'll explore
- writing more generic rules using wildcards;
- typing fewer filenames by using more templates;
- providing a list of default output files to produce;
- running commands in parallel on a single computer
- loading lists of filenames from spreadsheets
- configuring workflows with input files
Comments !