The week of February 21-25, 2022, we hosted the first Common Fund Data Ecosystem (CFDE) Hackathon. The goals of the hackathon were to increase familiarity with data sets from Common Fund programs and work towards cross-cutting, integrative analyses.

We invited members of the CFDE to propose hackathon sessions to introduce their Common Fund data sets and provide technical support while attendees explored the data. Sessions featured data from the CFDE Portal, Human Microbiome Project (HMP), Extracellular RNA Communication (exRNA), Metabolomics Workbench (MW), and Signature Commons Library of Integrated Network-Based Cellular Signatures (SigCom LINCS).
The Hackathon Sessions
All sessions were recorded and can be viewed on the Session Details and Recordings page of the hackathon website!
This virtual event began Monday morning with a welcome address by Dr. Titus Brown (UC Davis) followed by presentations from each Common Fund Program to give a brief overview of their data and session goals.
On Monday afternoon, Dr. Amanda Charbonneau (UC Davis) taught attendees how to use the CFDE Portal to find datasets from participating Common Fund programs. Dr. Charbonneau used HMP data as a motivating example, then helped attendees discover data sets from other programs. These datasets are quite large, so on Tuesday afternoon, Dr. Charbonneau taught a second session on how to download and process data from the CFDE Portal using Amazon Web Services (AWS). Attendees were provided with AWS accounts that they could use to analyze data discovered through the portal.
On Tuesday morning, Emily LaPlante and Keyang Yu (Baylor College of Medicine) provided an overview of the exRNA Atlas, which contains over 7,500 small RNA sequences and qPCR profiles from human and mouse, and introduced attendees to a variety of software tools for exploring RNA binding proteins. This session explored two use cases:
1) Finding RNA binding proteins and their associated RNA cargo in a variety of human biofluids and exploring their utility as biomarkers
2) Exploring other sites across the genome by intersecting exRNA Atlas data with regions of interest using BedGraph files, as well as applying this approach to other datasets.
On Wednesday morning, Eoin Fahy and Mano Maurya (UCSD) introduced the Metabolomics Workbench database which contains over 164,000 molecular structures covering 100+ species! Attendees learned how to interact with the Metabolomics Workbench Portal and then viewed a demonstration of MetENP, an R package that enables detection of significant metabolites from metabolite information.
The final data-driven hackathon session took place on Thursday afternoon. John Erol Evangelista (Mt Sinai) introduced the SigCom LINCS API which contains over 1.5 million gene expression signatures from LINCS, the Gene Expression Tissue Project (GTEx), and Gene Expression Omnibus databases (GEO). Then, Daniel Clarke (Mt Sinai) gave an introduction to building Appyters and how to use the SigCom LINCS APIs within Appyters.
On Friday we ran a Wrap Up and Future Directions session for presenters to recap what happened at their sessions and talk about future goals for their tools. This allowed everyone to learn about sessions they might not have attended, and possibly spark interest in watching the video recording of the session.
Reflection
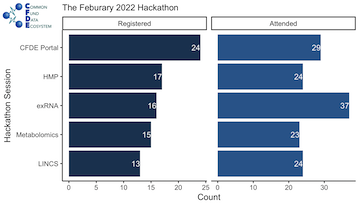
Overall the sessions were well attended and well received! In a pre-hackathon survey, we asked participants which hackathon sessions they were interested in attending. More people attended each session than we anticipated, which indicated that the introduction session on Monday was critical for spurring interest.
According to our survey, participants walked away with a greater understanding of Common Fund databases and tools, so we achieved our main goal of increasing familiarity with the diverse datasets supported by the Common Fund Data Ecosystem. Additionally, our team of trainers identified new Common Fund datasets that we plan on integrating into our training program in the future.
A common observation was that some sessions felt more like webinars or workshops than what the name "hackathon" implies. For our next hackathon, we will work with presenters to define sessions as webinars (a demonstration of a data tool), workshops (a training event with live coding), or hackathons (a defined problem that participants work on). We also received requests for more advanced notice and information about the content of sessions, which we will incorporate into our next round of event planning.
This event, along with many other online events, lacked the sense of community that can be present with in person multi-day events. We tried using GitHub Issues or Discussions to foster conversations between participants, but these tools were rarely used. We are thinking about how to address this for our next event, and we're open to feedback!
Finally, the hackathon coordination team would like to reiterate our thanks to all Common Fund groups that ran sessions for this event! We could not have achieved a diversity of datasets and tools at this event without your time and efforts.
Next steps
We are excited to announce that the second CFDE Hackathon will take place April 25 - 29th! We're going to fine tune the event with the feedback from our February event, and we hope you will join us!
If you are interested in learning more about attending the April 2022 Hackathon as a participant, please register here! We hope to see you there :)
The Common Fund Data Ecosystem Training Program is funded by the National Institutes of Health (1OT3OD025459-01).
Comments !